1. Introduction
Pharmacometrics underpins model-informed drug development through population PK/PD modeling, exposure–response analyses, and PBPK/QSP frameworks. Increasing dataset complexity, adaptive trial designs, and demand for faster timelines are reshaping expectations. AI and ML methods are gaining traction not as replacements, but as accelerators and enhancers.
This review presents recent (2022–2025) advancements in AI’s role across pharmacometric workflows, supported by detailed examples. It highlights where AI delivers immediate benefits—data QC, covariate discovery, biomarker integration, coding efficiency—and where exciting innovations lie on the horizon.
2. AI in Data Assembly and Quality Control
Creating analysis-ready datasets from multiple sources remains laborious. AI tools are now enhancing three key areas:
- Automated data extraction. NLP models mine study reports and registries to capture dosing schedules, sampling times, and biomarkers.
- Data cleaning and QC. Anomaly detection models identify implausible values or inconsistent units faster and more effectively than rule-based checks.
- Missing data imputation. Random forests and neural nets are being used to impute covariate gaps. A 2022 study found these ML-based methods performed comparably to predictive mean matching with greater flexibility in nonlinear scenarios [1].
Concrete examples:
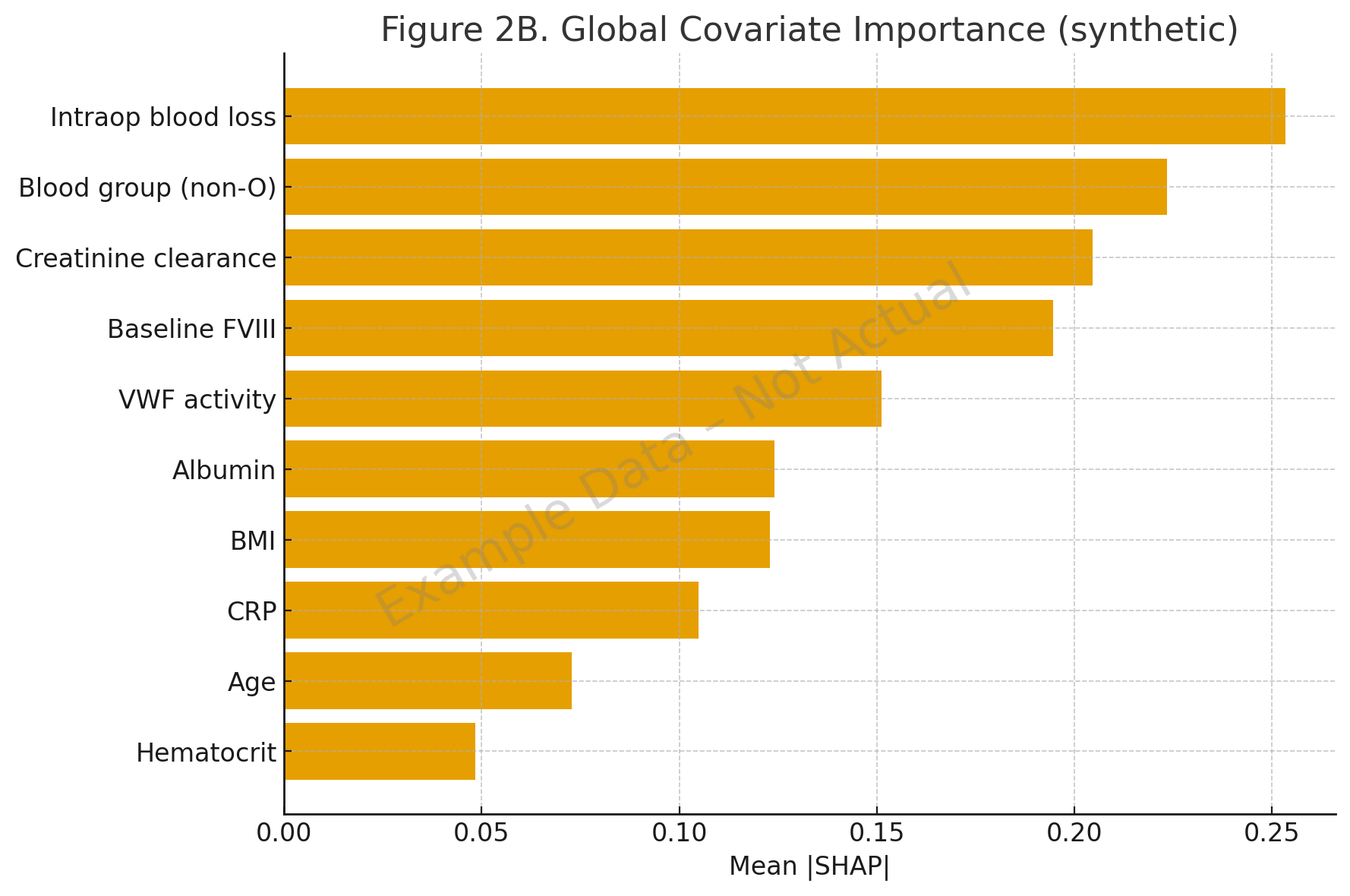
- A 2022 hemophilia A study applied SHAP with random forest and XGBoost to infer functional relationships between covariates and clearance. It confirmed known effects (blood group, baseline FVIII, VWF) and revealed nonlinear BMI interactions and synergistic effects of VWF with surgical blood loss [2].
- ML-based anomaly detection and imputation reduced QC cycle times, capturing errors traditional range checks missed.

3. AI in PK and PK–PD Modeling
Traditional pharmacometric approaches are robust but slow when exploring large model spaces. Several studies illustrate how AI is speeding up model development:
- qDarwin toolbox (2023). Combines ML algorithms with NONMEM to search globally across model structures. It reduces labor and increases objectivity while preserving interpretability [3].
- Symbolic regression for covariate structure (2024). Wahlquist and colleagues applied symbolic regression networks to a large propofol PK dataset (n = 1031). The method matched or improved fit, relied on fewer covariates, and eliminated manual model building [4].
| Year | Study | Method | Application | Key Finding |
|---|---|---|---|---|
| 2023 | qDarwin toolbox | ML integrated with NONMEM | Global model search | Reduced bias and runtime vs manual search [3] |
| 2024 | Wahlquist et al. | Symbolic regression | Propofol PK (n = 1031) | Accurate predictions with fewer covariates [4] |
| 2024 | Ridge regression prescreening | Regularized regression | Covariate screening vs SCM | Improved F1 (0.86 vs 0.50), runtime reduced > 40% [5] |
| 2025 | Shap-Cov workflow (Genentech) | Explainable ML (SHAP) | Covariate identification | Quantified influence with uncertainty estimates [6] |
4. Covariate Analysis and Biomarker Integration
Determining covariate effects and incorporating biomarkers remains one of the most subjective parts of modeling. AI is introducing rigor and automation.
4.1 Automated covariate screening
- Ridge regression vs SCM showed faster and more accurate prescreening in simulations [5].
- A 2025 systematic review summarized methods including AALASSO, genetic algorithms, random forests, and hybrid ML [7].
4.2 Machine learning with explainability (SHAP)
- Shap-Cov workflow (Genentech 2025) integrated SHAP-based explainability into covariate identification with uncertainty quantification and significance testing [6].
- In hemophilia A, SHAP revealed interactions like non-O blood group with low FVIII disproportionately increasing clearance, and nonlinear BMI effects [2].
4.3 Neural nets with stochastic gates
- Preprints (2025) describe neural nets with stochastic gates that learn sparse covariate sets under high correlation [8].
4.4 Biomarker integration
- A 2025 VAE + LASSO approach applied to tacrolimus PK uncovered key covariates (SNPs, albumin, hemoglobin) from high-dimensional biomarker profiles, achieving a MAPE of 2.26% and stable selection [9].

| Year | Study | Method | Application | Key Finding |
|---|---|---|---|---|
| 2022 | Hemophilia A SHAP study | RF/XGBoost + SHAP | Clearance & V1 covariates | Identified BMI nonlinearity, FVIII–blood group synergy [2] |
| 2024 | Ridge prescreening | Ridge regression | Covariate screening | Improved accuracy and faster runtime vs SCM [5] |
| 2025 | Shap-Cov workflow (Genentech) | SHAP with significance testing | PopPK covariate selection | Transparent ranking and uncertainty quantification [6] |
| 2025 | NN with stochastic gates | Sparse neural nets | Monalizumab dataset | Recovered expert-identified and new covariates [8] |
| 2025 | VAE + LASSO | Hybrid generative + regression | Tacrolimus PK covariates | MAPE 2.26%, robust biomarker selection [9] |
5. AI-Assisted Coding and Agentic Modeling
AI copilots are now used to generate initial NONMEM or R scripts from plain-text descriptions:
- A 2025 ArXiv study evaluated seven AI agents (including o1 and GPT-4.1) on 13 PK/PD modeling tasks. With optimized prompting, both achieved perfect accuracy, producing runnable NONMEM control streams [10].
- A 2024 evaluation of ChatGPT-4.0 and Gemini Ultra 1.0 showed that both could generate initial NONMEM templates but outputs contained syntax errors requiring manual correction [11].
Agentic AI systems are emerging:
- A 2025 review in Clinical and Translational Science described workflows where specialized agents manage data extraction, model evaluation, and simulation with human oversight [12].
| Year | Study | Method | Application | Key Finding |
|---|---|---|---|---|
| 2024 | PubMed 38656706 | ChatGPT-4.0, Gemini | NONMEM code generation | Produced drafts but required manual correction [11] |
| 2025 | ArXiv 2507.08144 | GPT-4.1, o1 | NONMEM tasks (13 models) | 100% accuracy with optimized prompts [10] |
| 2025 | PubMed 40055986 | Agentic workflows | Multi-agent orchestration | Automated modeling loops with human oversight [12] |
6. Workflow Acceleration and Integration
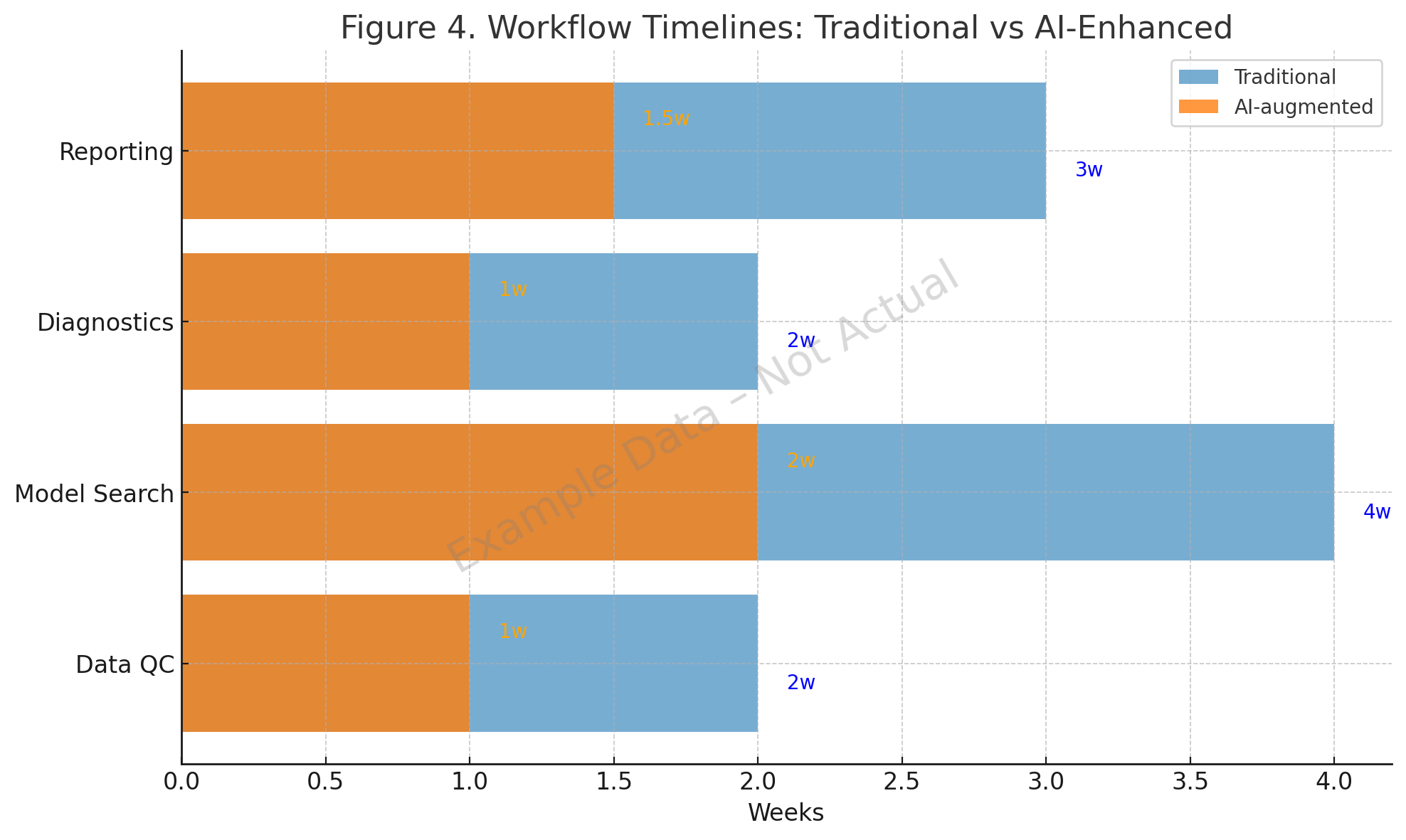
Concrete studies show that AI reduces pharmacometric cycle times:
- Data QC. SHAP-based hemophilia A analysis (2022) flagged nonlinear covariate interactions [2]. ML imputation methods provided robust handling of missing PK samples [1].
- Model search. The qDarwin toolbox (2023) and symbolic regression (2024) improved search efficiency [3,4].
- Diagnostics and reporting. The Pharmpy AMD module (2024) generated diagnostics reliably [5], and AI copilots in 2025 embedded diagnostic templates into generated NONMEM code [10].


7. Regulatory and Practical Considerations
- Transparency. Black-box models must be supplemented with explainability tools.
- Validation. AI-generated code and datasets require documented audit trails.
- Regulatory perspectives. FDA and EMA have encouraged risk-based validation and context-of-use definitions for AI in drug development.
AI should be viewed as an assistant. Analysts remain accountable for interpretation and scientific rigor.
8. Emerging and Future Directions
- Real-time model updating during ongoing trials.
- Generative synthetic cohorts to stress-test trial robustness.
- Automated pharmacometric reporting in CSRs and regulatory submissions.
- Cloud-native agentic platforms scaling overnight model exploration.
- Multi-omic and digital biomarker integration into PK/PD frameworks.
9. Conclusion
AI is already easing pain points in pharmacometric workflows. Examples such as ridge regression prescreening, symbolic regression for propofol PK, SHAP-Cov workflows, stochastic-gate neural nets, and VAE-LASSO covariate discovery demonstrate tangible advances. Similarly, copilots for NONMEM coding and agentic frameworks for model orchestration highlight the evolving toolkit.
The principle remains balance. Mechanistic expertise guides model design. AI improves efficiency, scale, and reproducibility. Pragmatic adoption will allow pharmacometric teams—especially in small and mid-size biotech—to deliver analyses that are faster, richer, and no less rigorous than before.
References
- Antibiotics. 2024;13(12):1203. Machine learning-based imputation in pharmacometric datasets.
- Holford et al. SHAP analysis in hemophilia A population PK. Front Pharmacol. 2022; PMC9381890.
- qDarwin toolbox. Automated model selection with NONMEM. CPT Pharmacometrics Syst Pharmacol. 2023.
- Wahlquist et al. Symbolic regression for PK covariate modeling. J Pharmacokinet Pharmacodyn. 2024.
- Ridge regression prescreening for covariates. Pharm Res. 2024.
- Zhang et al. Shap-Cov workflow. CPT Pharmacometrics Syst Pharmacol. 2025.
- Systematic review of covariate selection methods. Clin Pharmacokinet. 2025.
- Neural nets with stochastic gates for covariate selection. bioRxiv. 2025.
- Tacrolimus covariate analysis with VAE + LASSO. arXiv preprint. 2025.
- Evaluation of AI copilots for NONMEM tasks. arXiv preprint. 2025;2507.08144.
- Comparative study of ChatGPT and Gemini for NONMEM coding. Br J Clin Pharmacol. 2024; PubMed ID 38656706.
- Agents for Change: Agentic AI in Clinical Pharmacology. Clin Transl Sci. 2025; PubMed ID 40055986.
Abbreviations
- AI: Artificial Intelligence
- AMD: Automatic Model Development
- BIC: Bayesian Information Criterion
- BMI: Body Mass Index
- CL: Clearance
- CrCl: Creatinine Clearance
- CRP: C-reactive Protein
- CSR: Clinical Study Report
- GAN: Generative Adversarial Network
- ML: Machine Learning
- MAPE: Mean Absolute Percentage Error
- NONMEM: Nonlinear Mixed Effects Modeling
- PBPK: Physiologically Based Pharmacokinetic
- PD: Pharmacodynamic
- PK: Pharmacokinetic
- PopPK: Population Pharmacokinetics
- QSP: Quantitative Systems Pharmacology
- RF: Random Forest
- SCM: Stepwise Covariate Modeling
- SHAP: Shapley Additive Explanations
- SNP: Single Nucleotide Polymorphism
- VAE: Variational Autoencoder
- VPC: Visual Predictive Check
- VWF: von Willebrand Factor
