Module 2.6.4 is the reviewer’s fast path to your nonclinical ADME story. Done well, it shows what was studied, what was learned, and why those data are sufficient for the current stage of development and the proposed clinical plan, with traceability back to Module 4 and forward to the clinical overview (2.7.1) [1].
What belongs in Module 2.6.4
- Absorption: solubility, permeability, oral bioavailability, and any food-effect rationale carried into 2.7.1 [1][7][6].
- Distribution: protein binding, tissue distribution (including QWBA where available), CNS/blood-brain barrier, TMDD considerations for biologics [1].
- Metabolism: in-vitro CLint (hepatocytes/HLM), major pathways (e.g., CYP3A4, UGT1A1), metabolite ID, and human relevance; link to DDI strategy [1][5][12].
- Excretion: mass balance (if available), urine/feces/bile split, and any renal/hepatic impairment considerations that inform the clinical plan [1].
- Bioanalysis & Assay Validation: assay type and selectivity, accuracy/precision, stability, and a pointer to validation reports [3][4][13].

Modality-specific expectations
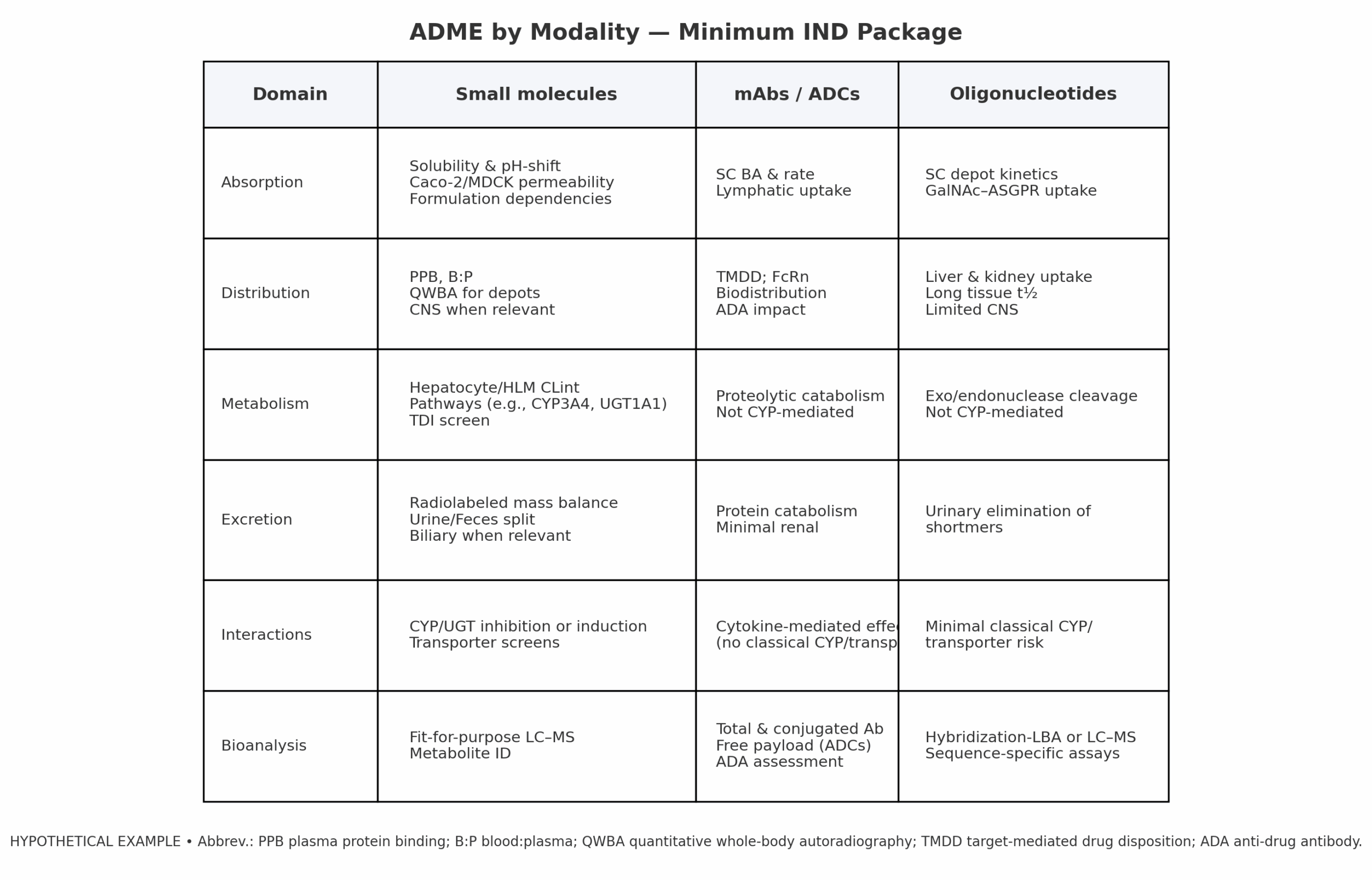
Regulatory expectations vary by modality. Your tables should make these differences obvious so reviewers can quickly judge adequacy [1].
- Small molecules: Oral feasibility and dissolution rationale; permeability/efflux profile; enzyme/transporters (inhibition, induction, phenotyping); mass balance and metabolite ID where available; a DDI plan or PBPK justification carried into clinical [5][6].
- mAbs / ADCs: Biodistribution and TMDD considerations; FcRn engagement; immunogenicity (ADA) impact on PK; for ADCs, total and conjugated antibody with free-payload tracking and linker stability [1].
- Oligonucleotides: Uptake and tissue retention (e.g., liver/kidney), sequence-specific bioanalytical methods, and limited interaction risk via classical CYP/transporters [1].
Applied examples (brief case snapshots)
Small molecules — CYP3A risk translated into clinical strategy
EGFR inhibitor programs commonly evaluate CYP3A liability early. For example, published work with osimertinib shows that strong CYP3A inducers can substantially reduce exposure and should be avoided, while strong inhibitors have a smaller effect; labels for this class typically incorporate dose-management or avoidance language based on that risk profile [8]. In 2.6.4, summarize the in-vitro phenotyping/inhibition/induction and any PBPK, then link to 2.7.1 (planned clinical DDI with a sensitive substrate or management via labeling) [5].
Biologics — disease-DDI via cytokine normalization
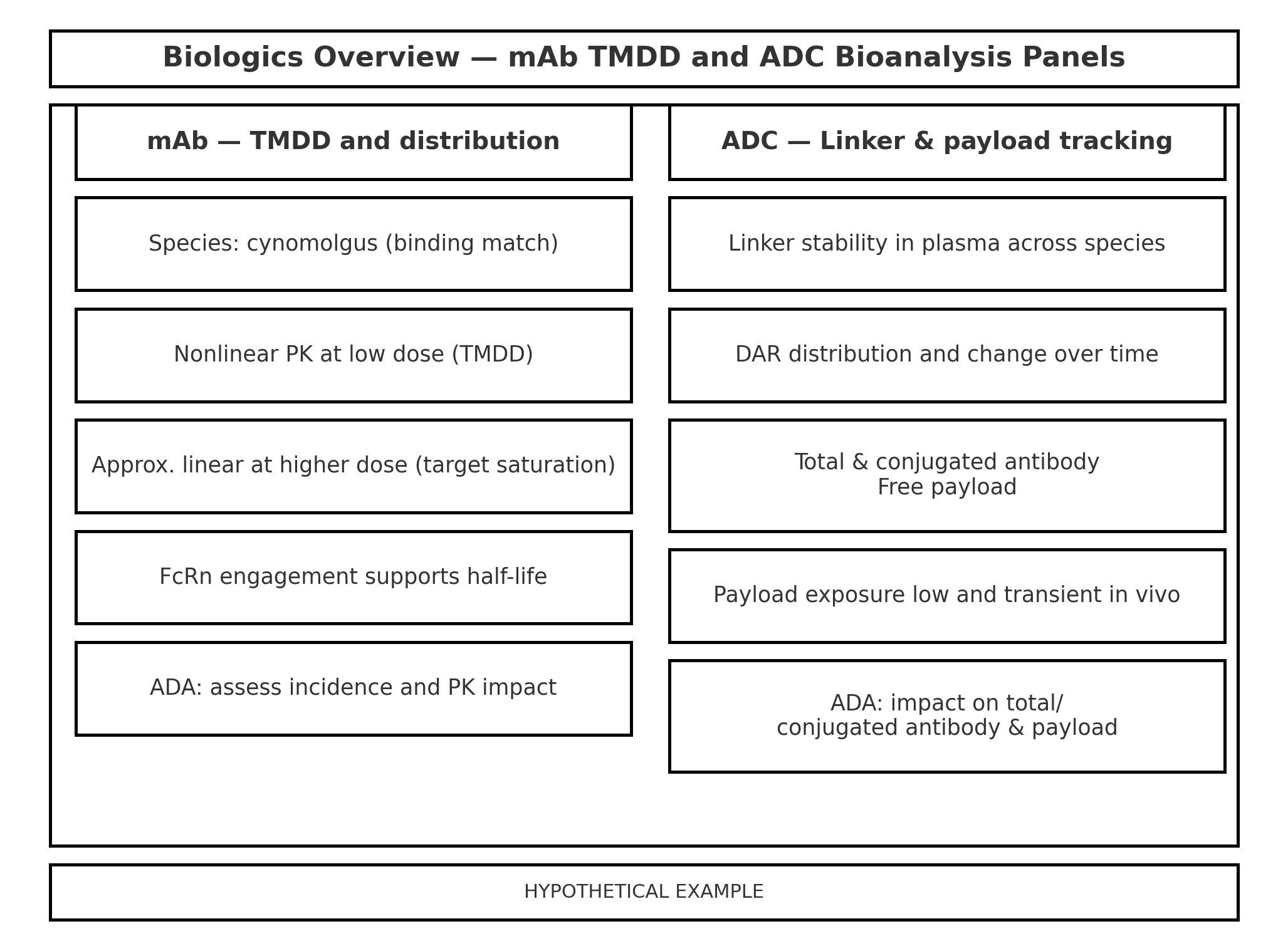
Monoclonal antibodies generally do not cause classical CYP-mediated DDIs. However, IL-6 blockade can normalize CYP3A4 activity; with tocilizumab, simvastatin exposure decreased after treatment because inflammation-driven CYP suppression was reversed. This is a classic “disease-DDI” scenario to acknowledge in 2.6.4 and carry forward into clinical labeling considerations [9][16].
ADCs — catabolism, linker stability, and payload tracking
For ado-trastuzumab emtansine (T-DM1), the nonclinical/clinical PK package emphasizes catabolism to DM1 rather than CYP metabolism, minimal renal excretion of intact conjugate, and the need to track total/conjugated antibody and free-payload exposures. In 2.6.4, keep the narrative concise and point to Module 4 for assay details and exposure relationships [10][14].
Oligonucleotides — targeted hepatic uptake with limited classical DDI risk
Inclisiran (siRNA) illustrates GalNAc-mediated hepatocyte uptake, long tissue residence, and limited potential for classical CYP/transporter interactions. Regulatory reviews and product information highlight minimal interaction liability and sequence-specific bioanalysis. Reflect that pattern in 2.6.4 and link to Module 5 for assay validation [11][15][7].
Sample authoring language you can adapt
(Fictional asset: CPDS-1001)
- Absorption (summary line): “CPDS-1001 shows high permeability in Caco-2 and no clinically meaningful efflux liability; the fed-state formulation is selected to mitigate a pH-dependent solubility drop and is consistent with food-effect planning in 2.7.1 [1][7].”
- Metabolism: “Human hepatocytes/HLM indicate oxidative clearance via CYP3A4 and minor glucuronidation (UGT1A1). Inhibition/induction screens and PBPK underpin the clinical DDI plan in 2.7.1 [5][12].”
- Excretion: “Available data support dual renal/hepatic elimination; full radiolabeled mass balance is planned at Phase 2/3 transition; current risk is acceptable for FIH with monitoring as outlined in the protocol [1].”
- Bioanalysis: “Primary LC–MS/MS method met acceptance criteria (selectivity, accuracy, precision, stability) with ISR >67%; validation report cross-referenced in Module 5 [3][4][13].”
Traceability to Module 4 and to clinical
- Every conclusion in 2.6.4 maps to a specific Module 4 report, table, or figure (study ID, table, page).
- Forward links: pull through what matters for 2.7.1 (food-effect rationale, DDI risk/PBPK, variability that impacts dose selection).
- Label implications: if anything in ADME could influence labeling later (e.g., strong CYP3A4 interaction or clinically meaningful food effect), state it plainly and flag how it will be addressed [5].
Common pitfalls and reviewer questions
- Tables list many studies but the narrative lacks the “so what” that changes the program’s risk posture.
- Bioanalytical method summary omits key performance characteristics (selectivity, stability) or the pointer to validation [3][4][13].
- Transporter or enzyme work is mismatched to the clinical route or expected exposures [5].
- Insufficient linkage to 2.7.1 and to the clinical pharmacology/DDI plan.
References
- ICH M4S(R2): Nonclinical Overview and Nonclinical Summaries (Module 2.4/2.6).
- EMA copy of ICH M4S (Module 2.6 organization).
- FDA (2018) Bioanalytical Method Validation.
- EMA Bioanalytical Method Validation Guideline.
- FDA (2020) Clinical Drug Interaction Studies — CYP/Transporter-Mediated.
- ICH M9 (2019) BCS-Based Biowaivers.
- FDA Food-Effect Bioavailability and Fed BE Studies.
- Vishwanathan et al. (2018) Osimertinib: Impact of CYP3A modulators.
- Schmitt et al. (2011) Tocilizumab disease-DDI (simvastatin).
- Kadcyla (ado-trastuzumab emtansine) U.S. label — Clinical Pharmacology.
- Inclisiran (Leqvio) U.S. label — Clinical Pharmacology.
- FDA (2020) In Vitro Drug Interaction Studies (CYP/Transporters).
- ICH M10 (2023) Bioanalytical Method Validation and Study Sample Analysis.
- Tocilizumab label — CYP3A4 normalization note.
Abbreviations
ADA: anti-drug antibody
ADME: absorption, distribution, metabolism, excretion
BA: bioavailability
B:P: blood:plasma ratio (blood-to-plasma)
BCS: biopharmaceutics classification system
CLint: intrinsic clearance
CNS: central nervous system
CTD: Common Technical Document
DDI: drug–drug interaction
HLM: human liver microsomes
ISR: incurred sample reanalysis
PBPK: physiologically based pharmacokinetic
PK: pharmacokinetic
QWBA: quantitative whole-body autoradiography
TMDD: target-mediated drug disposition
